-
Abstract: Primary liver cancer is the most common type of malignant tumors in China. Due to the insufficient early diagnosis, most of the patients with liver cancer are diagnosed in the advanced stages, resulting in poor prognosis. Currently, the diagnostic methods for liver cancer are dependent on the traditional biomarkers alpha-fetoprotein (AFP), together with ultrasound and other imaging techniques. However, the sensitivity and specificity of these biomarkers are not optimal. Noninvasive, serum-based biomarkers are urgently required for the early detection of HCC.This review briefly describes the progresses in the potential biomarkers for HCC diagnosis.
-
Key words:
- hepatocellular carcinoma /
- biomarkers /
- research progress
-
-
[1] Sung H, Ferlay J, Siegel RL, et al. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries[J]. CA Cancer J Clin, 2021, 71(3): 209-249. doi: 10.3322/caac.21660
[2] 国家卫生健康委办公厅. 原发性肝癌诊疗指南(2022年版)[J]. 临床肝胆病杂志, 2022, 38(2): 288-303. doi: 10.3969/j.issn.1001-5256.2022.02.009
[3] Siegel RL, Miller KD, Jemal A. Cancer statistics 2020[J]. CA Cancer J Clin, 2020, 70(1): 7-30. doi: 10.3322/caac.21590
[4] 陆荫英, 赵海涛, 程家敏, 等. 肝胆肿瘤分子诊断临床应用专家共识[J]. 临床肝胆病杂志, 2020, 36(7): 1482-1488. doi: 10.3969/j.issn.1001-5256.2020.07.008
[5] Zhang R, Lin HM, Broering R, et al. Dickkopf-1 contributes to hepatocellular carcinoma tumorigenesis by activating the Wnt/β-catenin signaling pathway[J]. Signal Transduct Target Ther, 2019, 4: 54. doi: 10.1038/s41392-019-0082-5
[6] Niu J, Li W, Liang C, et al. EGF promotes DKK1 transcription in hepatocellular carcinoma by enhancing the phosphorylation and acetylation of histone H3[J]. Science Signaling, 2020, 13(657): eabb5727. doi: 10.1126/scisignal.abb5727
[7] 安琳, 袁方, 覃文新, 等. 磁微粒化学发光法测定DKK-1在肝细胞癌诊断中的价值[J]. 国际检验医学杂志, 2017, 38(13): 1729-1731. doi: 10.3969/j.issn.1673-4130.2017.13.001
[8] 吴晓霞. DKK1、CTCs在原发性肝癌TACE术前后表达变化及预后价值研究[D]. 苏州大学, 2018.
[9] Shen Q, Fan J, Yang XR, et al. Serum DKK1 as a protein biomarker for the diagnosis of hepatocellular carcinoma: a large-scale, multicentre study[J]. Lancet Oncol, 2012, 13(8): 817-826. doi: 10.1016/S1470-2045(12)70233-4
[10] Yap L, Tay HG, Nguyen MTX, et al. Laminins in Cellular Differentiation[J]. Trends Cell Biology, 2019, 29(12): 987-1000. doi: 10.1016/j.tcb.2019.10.001
[11] Yasuda H, Nakagawa M, Kiyokawa H, et al. Unique Biological Activity and Potential Role of Monomeric Laminin-γ2 as a Novel Biomarker for Hepatocellular Carcinoma: A Review[J]. Int J Mol Sci, 2019, 20(1): 226. doi: 10.3390/ijms20010226
[12] Govaere O, Wouters J, Petz M, et al. Laminin-332 sustains chemoresistance and quiescence as part of the human hepatic cancer stem cell niche[J]. J Hepatol, 2016, 64(3): 609-617. doi: 10.1016/j.jhep.2015.11.011
[13] Kiyokawa H, Yasuda H, Oikawa R, et al. Serum monomeric laminin-γ2 as a novel biomarker for hepatocellular carcinoma[J]. Cancer Sci, 2017, 108(7): 1432-1439. doi: 10.1111/cas.13261
[14] Zhang S, Shao J, Su F. Prognostic significance of STIP1 expression in human cancer: A meta-analysis[J]. Clin Chim Acta, 2018, 486: 168-176. doi: 10.1016/j.cca.2018.07.037
[15] Krafft U, Tschirdewahn S, Hess J, et al. STIP1 Tissue Expression Is Associated with Survival in Chemotherapy-Treated Bladder Cancer Patients[J]. Pathol Oncol Res, 2020, 26(2): 1243-1249. doi: 10.1007/s12253-019-00689-y
[16] Wang HS, Tsai CL, Chang PY, et al. Positive associations between upregulated levels of stress-induced phosphoprotein 1 and matrix metalloproteinase-9 in endometriosis/adenomyosis[J]. PLoS One, 2018, 13(1): e0190573. doi: 10.1371/journal.pone.0190573
[17] Chen Z, Xu L, Su T, et al. Autocrine STIP1 signaling promotes tumor growth and is associated with disease outcome in hepatocellular carcinoma[J]. Biochem Biophys Res Commun, 2017, 493(1): 365-372. doi: 10.1016/j.bbrc.2017.09.016
[18] Ma XL, Tang WG, Yang MJ, et al. Serum STIP1, a Novel Indicator for Microvascular Invasion, Predicts Outcomes and Treatment Response in Hepatocellular Carcinoma[J]. Front Oncol, 2020, 10: 511. doi: 10.3389/fonc.2020.00511
[19] Somm E, Jornayvaz FR. Fibroblast Growth Factor 15/19: From Basic Functions to Therapeutic Perspectives[J]. Endocr Rev, 2018, 39(6): 960-989. doi: 10.1210/er.2018-00134
[20] Maeda T, Kanzaki H, Chiba T, et al. Serum fibroblast growth factor 19 serves as a potential novel biomarker for hepatocellular carcinoma[J]. BMC Cancer, 2019, 19(1): 1088. doi: 10.1186/s12885-019-6322-9
[21] Prendergast GC, Mondal A, Dey S, et al. nflammatory Reprogramming with IDO1 Inhibitors: Turning Immunologically Unresponsive'Cold' Tumors'Hot'[J]. Trends Cancer, 2018, 4(1): 38-58. doi: 10.1016/j.trecan.2017.11.005
[22] Yuhei S, Takeshi H, Junji N, et al. The Role of Indoleamine 2, 3-Dioxygenase in Diethylnitrosamine-Induced Liver Carcinogenesis[J]. PLos One, 2016, 11(1): e0146279. doi: 10.1371/journal.pone.0146279
[23] Stepien M, Duarte-Salles T, Fedirko V, et al. Alteration of amino acid and biogenic amine metabolism in hepatobiliary cancers: Findings from a prospective cohort study[J]. Int J Cancer, 2016, 138(2): 348-360. doi: 10.1002/ijc.29718
[24] Bekki S, Hashimoto S, Yamasaki K, et al. Serum kynurenine levels are a novel biomarker to predict the prognosis of patients with hepatocellular carcinoma[J]. PLoS one, 2020, 15(10): e0241002. doi: 10.1371/journal.pone.0241002
[25] Zhang L, Tian R, Yao X, et al. Milk Fat Globule-EGF Factor 8 improves Hepatic Steatosis and Inflammation[J]. Hepatology, 2020, 73(2): 586-605.
[26] Shimagaki T, Yoshio S, Kawai H, et al. Serum milk fat globule-EGF factor 8(MFG-E8) as a diagnostic and prognostic biomarker in patients with hepatocellular carcinoma[J]. Sci Rep, 2019, 9(1): 15788. doi: 10.1038/s41598-019-52356-6
[27] Shimizu Y, Suzuki T, Yoshikawa T, et al. Next-Generation Cancer Immunotherapy Targeting Glypican-3[J]. Front oncol, 2019, 9: 248. doi: 10.3389/fonc.2019.00248
[28] Li D, Li N, Zhang YF, et al. Persistent Polyfunctional Chimeric Antigen Receptor T Cells That Target Glypican 3 Eliminate Orthotopic Hepatocellular Carcinomas in Mice[J]. Gastroenterology, 2020, 158(8): 2250-2265. doi: 10.1053/j.gastro.2020.02.011
[29] Sun B, Huang Z, Wang B, et al. Significance of Glypican-3(GPC3) Expression in Hepatocellular Cancer Diagnosis[J]. Med Sci Monit, 2017, 23: 850-855. doi: 10.12659/MSM.899198
[30] 谢春梅, 徐伟文. GPC3在肝癌诊治中的价值及存在问题[J]. 分子诊断与治疗杂志, 2016, 8(1): 1-6. doi: 10.3969/j.issn.1674-6929.2016.01.001
[31] Jeon Y, Jang ES, Choi YS, et al. Glypican-3 level assessed by the enzyme-linked immunosorbent assay is inferior to alpha-fetoprotein level for hepatocellular carcinoma diagnosis[J]. Clin Mol Hepatol, 2016, 22(3): 359-365. doi: 10.3350/cmh.2016.0033
[32] Nomair AM, Issa NM, Madkour MA, et al. The clinical significance of serum miRNA-224 expression in hepatocellular carcinoma[J]. Clin Exp Hepatol, 2020, 6(1): 20-27. doi: 10.5114/ceh.2020.93052
[33] Shigeyasu K, Toden S, Zumwalt TJ, et al. Emerging Role of MicroRNAs as Liquid Biopsy Biomarkers in Gastrointestinal Cancers[J]. Clin Cancer Res, 2017, 23(10): 2391-2399. doi: 10.1158/1078-0432.CCR-16-1676
[34] An Y, Gao S, Zhao WC, et al. Novel serum microRNAs panel on the diagnostic and prognostic implications of hepatocellular carcinoma[J]. World J Gastroenterol, 2018, 24(24): 2596-2604. doi: 10.3748/wjg.v24.i24.2596
[35] Liu Y, Zhang Y, Wu H, et al. miR-10a suppresses colorectal cancer metastasis by modulating the epithelial-to-mesenchymal transition and anoikis[J]. Cell Death Dis, 2017, 8(4): e2739. doi: 10.1038/cddis.2017.61
[36] Amr KS, Elmawgoud Atia HA, Elazeem Elbnhawy RA, et al. Early diagnostic evaluation of miR-122 and miR-224 as biomarkers for hepatocellular carcinoma[J]. Genes Dis, 2017, 4(4): 215-221. doi: 10.1016/j.gendis.2017.10.003
[37] Li C, Deng M, Hu J, et al. Chronic inflammation contributes to the development of hepatocellular carcinoma by decreasing miR-122 levels[J]. Oncotarget, 2016, 7(13): 17021-17034. doi: 10.18632/oncotarget.7740
[38] 刘斌, 汤晓莉. 乙型病毒性肝炎患者血小板miRNA的表达水平及临床意义[J]. 临床血液学杂志, 2020, 33(4): 233-236. https://www.cnki.com.cn/Article/CJFDTOTAL-LCXZ202004001.htm
[39] Huang Z, Zhou JK, Peng Y, et al. The role of long noncoding RNAs in hepatocellular carcinoma[J]. Mol Cancer, 2020, 19(1): 77. doi: 10.1186/s12943-020-01188-4
[40] Kamel MM, Matboli M, Sallam M, et al. Investigation of long noncoding RNAs expression profile as potential serum biomarkers in patients with hepatocellular carcinoma[J]. Transl Res, 2016, 168: 134-145. doi: 10.1016/j.trsl.2015.10.002
[41] Szabo L, Salzman J. Detecting circular RNAs: bioinformatic and experimental challenges[J]. Nat Rev Genet, 2016, 17(11): 679-692.
[42] Wang S, Zhang K, Tan S, et al. Circular RNAs in body fluids as cancer biomarkers: the new frontier of liquid biopsies[J]. Mol Cancer, 2021, 20(1): 13. doi: 10.1186/s12943-020-01298-z
[43] Zhang X, Zhou H, Jing W, et al. The Circular RNA hsa_circ_0001445 Regulates the Proliferation and Migration of Hepatocellular Carcinoma and May Serve as a Diagnostic Biomarker[J]. Dis Markers, 2018, 2018: 3073467.
[44] Sun XH, Wang YT, Li GF, et al. Serum-derived three-circRNA signature as a diagnostic biomarker for hepatocellular carcinoma[J]. Cancer Cell Int, 2020, 20: 226. doi: 10.1186/s12935-020-01302-y
[45] Ye Q, Ling S, Zheng S, et al. Liquid biopsy in hepatocellular carcinoma: circulating tumor cells and circulating tumor DNA[J]. Mol Cancer, 2019, 18(1): 114. doi: 10.1186/s12943-019-1043-x
[46] Ahn JC, Teng PC, Chen PJ, et al. Detection of Circulating Tumor Cells and Their Implications as a Biomarker for Diagnosis, Prognostication, and Therapeutic Monitoring in Hepatocellular Carcinoma[J]. Hepatology, 2021, 73(1): 422-436. doi: 10.1002/hep.31165
-
| 引用本文: | 张丽芳, 袁野, 史林, 等. 新型肝癌标志物的临床应用研究进展[J]. 临床血液学杂志, 2022, 35(4): 297-302. doi: 10.13201/j.issn.1004-2806.2022.04.016 |
| Citation: | ZHANG Lifang, YUAN Ye, SHI Lin, et al. Advances in clinical application of novel liver cancer markers[J]. J Clin Hematol, 2022, 35(4): 297-302. doi: 10.13201/j.issn.1004-2806.2022.04.016 |
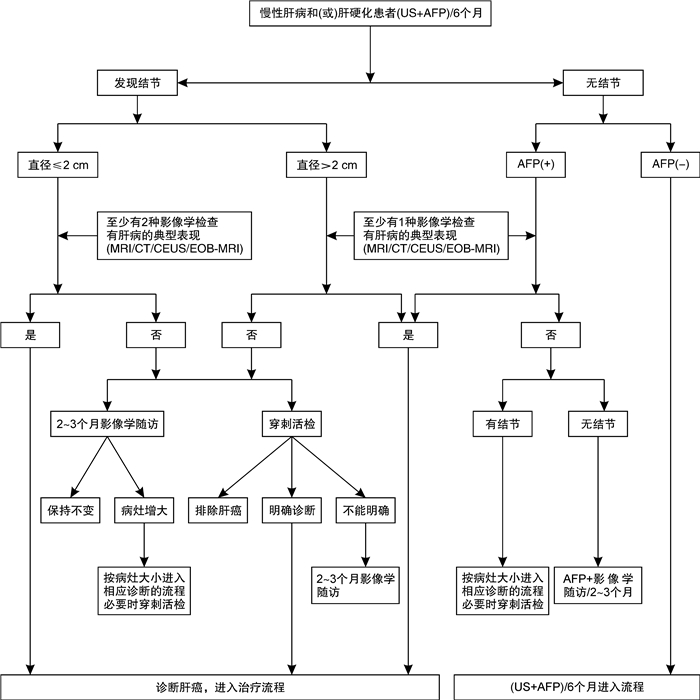
- Figure 1.


 下载:
下载: